Insertion
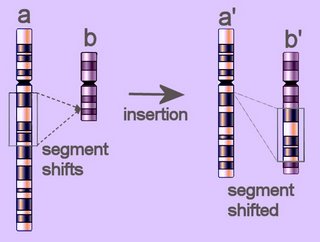
Diagram of insertion of a segment of chromosome a into chromosome b, resulting in shortened chromosome a' and lengthened chromosome b' (click to enlarge image).
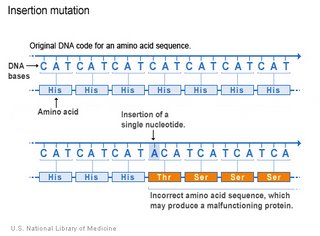
If the number of inserted bases is not a multiple of 3, insertion will cause frameshift, with serious consequences. A number of diseases are caused by insertions without frameshift - Huntington's chorea, Myotonic Dystrophy, Fragile X site A, Fragile X site E, Fragile X site F, Kennedy disease, SCA1, DRPLA.
Proteins gradually evolve by the accumulation of mutations.
Mobile elements called insertion sequences exist in nature. These sequences encode only the information necessary for their insertion into DNA. Depending upon the particular insertion sequence, they can insert at specific regions or at random.
Tables Mechanisms of Biological Evolution Gene Regulation in E.coli :
It has been known since the beginning of the twentieth century that unstable or variable gene loci occur in plants. Breakage-fusion (reunion)-bridges are formed during anaphase whenever two chromosomes fuse at their ends, generating a fusion product with two centromers. If these two fused chromosomes are subsequently carried to different poles than the regular chromosomes, a chromosomal fraction results. During the subsequent S-phase, the chromatid with a fused-fraction at its terminus will replicate, leading again to a fusion of the homologous chromatids. Consequently a chromosome comprising one chromatid with two centromeres will occur in the subsequent mitosis, rather than a chromosome from two chromatids and one centromere. Consequently, a second fraction occurs during anaphase when the second round of the cycle starts.
B. McClintock recognized (between ‘47 and ‘51) that the chromosomal fraction is restricted to certain sections of the chromosome, which she termed Ds (dissociation). The Ds segment is a mutator gene, which behaves like a pseudoallele that can be located at different gene loci. This mutator gene can insert itself into other genes, rendering them inactive. Thus, it is a control element that changes its location within the chromosome, causing mutations wherever it inserts. Such mutator genes are also called "jumping genes".
A further set of elements, the Ac (activation) elements, support the chromosomal fraction or a translocation of a Ds element. An Ac element can be regarded as a multiple allele, and it may occur different sites in all chromosomes. A number of gene loci are known to be influenced by the Ds-Ac-system or other control elements. Detection of the spm-system (suppressor-mutator) and the elucidation of its function established that the control elements not only act as switches (a yes/ no decision) but that they also modulate the degree of gene expression.
Insertion elements and transposons were first detected in bacterial DNA during the late sixties. This discovery explained the connection between transposons and the chromosome fraction control elements.
Table Mechanisms of Biological Evolution : Gene Regulation in E.coli :
External : Transposons part 1, transposons part 2 : Barbara McClintock and mobile genetic elements :
Labels: Barbara McClintock, deletion, insertion, MGE, mobile genetic elements, mutation, transposons









































Blue terms hyperlink to explanatory items. Linked items can also be found by way of the 'Links to this post' list at the base of some posts (once Blogger catches up!). Use the "back" function to return to the departure item.
Items occur within Sections. When visiting an item, the site title changes to purple – click on the title or “Home” to return to the main page. Topics are listed in the Site Map (click on arrow at top of sidebar). The site is searchable – once Blogger catches up – by way of the 'Search this blog' window at upper left.
When the number before the “Guide-Glossary” link (below each item) is greater than 0, the link provides a glossary of terms. Displayed as a pop-up when reading within a Section, or as sub-script when visiting an Item.
Allopatric speciation occurs when a geographical barrier sub-divides a parent species, resulting in geographic and reproductive isolation such that the descendent species can no longer interbreed upon removal of the barrier.
Anagenesis differs from cladogenesis in that one species progressively transforms into a replacement species when sufficient gene mutations fix in the descendant population. At this point, the ancestral species has become extinct. This mechanism is distinct from the increase in numbers of species generated by cladogenetic branching events.
Cladogenesis is the mechanism of speciation in which one or more lineages (clades) arise from an ancestral line. Such speciation events increase the variety of plants or animals through branching of the phylogenetic tree. Cladogenesis is differentiated from anagenesis, which is the in toto replacement of one species by an anatomically distinct species.
Monophyletic taxon or clade: an accurate grouping of only (opp. polyphyletic) and all (opp. paraphyletic) descendents of a shared common ancestor. A monopyletic group is genetically homogeneous and reflects evolutionary relationships.
Paraphyletic taxon or clade: a monophyletic group that excludes one or more discrete groups descended from the most recent common ancestral species of the entire group. Other descendent species of the most recent common ancestor have been excluded from the paraphyletic taxon, usually because of morphologic distinctiveness.
Phenetic system: groupings of organisms based on mutual similarity of phenotypic (physical and chemical) characteristics. Phenetic groupings may or may not correlate with evolutionary relationships.
Phylogenetic system: groups organisms based on shared evolutionary heritage. DNA and RNA sequencing techniques are considered to give the most meaningful phylogenies.
Phylogenetic separation into evolutionary relationships (clades), based on comparison of genomes is likely to supplant phenotypical (phenetic) taxonomies of the prokaryotes.
Peripatry (paripatry) is a subset of allopatry in which an isolated group has a smaller population than the parent group. Ernst Mayr introduced the term. Peripatric speciation occurs when the smaller sub-group of a species enters a novel niche within the range of the parent species, becoming geographically and reproductively isolated. Peripatric speciation (paripatric) is distinguished from allopatric speciation by the smaller size of the isolate group, and from sympatric speciation, which involves no barrier to breeding.
Polyphyletic taxon: opposite to monophyletic taxon: A polyphyletic group is mistakenly or improperly erected on the basis of homoplasy.—characteristics that have arisen despite not sharing a common ancestor. Homoplasy arises because of convergent evolution, parallelism, evolutionary reversals, horizontal gene transfer, or gene duplications. Polyphyletic taxa are genetically heterogeneous because members do not share a common ancestor.
Neontology is a branch of biology that emphasizes the study of modern biota (living or recent organisms) rather than fossilized organisms (paleontology).
Numerical Taxonomies are a common approach to phenetic taxonomy that employ a number of phenotypic characteristics to generate similarity coefficients that may be mapped in dendrograms. Groupings based on numerical taxonomy may or may not correlate with evolutionary relationships.
Taxonomies aim to group organisms according to shared characteristics against the background of biological diversity.
Sympatry involves no geographical separation of sub-populations of individuals. Sympatric speciation events occur most often in plants by the mechanism of polyploidy in which the number of chromosomes is doubled or tripled. John Maynard Smith proposed a model called disruptive speciation, in which homozygotes might have greater fitness than heterozygotes under some environmental conditions.