Conjugation
Conjugation is often likened to a form of sexual reproduction or mating, though it is merely a process by which donor bacteria deliver genetic material to recipient bacteria, utilizing tube-like connections called pili.
Bacteria often contain small circular, double-stranded DNA molecules that are termed plasmids. Bacterial plasmids are not connected to the main bacterial chromosomes and replicate independently. The donor bacterium contains conjugative or mobilizable genetic elements, usually a conjugative plasmid or episome plasmid that can integrate itself into the bacterial chromosome by genetic recombination. One such conjugative plasmid is called the F-plasmid. This is an episome about 100 thousand base-pairs in length, which carries its own origin of replication, called oriV. Most conjugative plasmids have systems ensuring that the recipient cell does not already contain a similar element, ensuring that there is only one copy of the F-plasmid in the F-positive bacterium. diagram of conjugation events diagram of molecular events
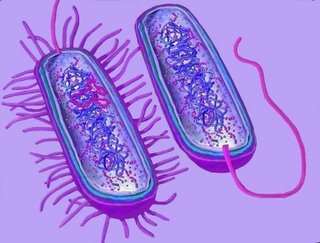
One cell contains an F-plasmid (pink), distinct from the prokaryotic genomer (blue). The cell with an F-plasmid also possesses pili, which make contact with the F-negative cell, which does not have pili (top right diagram).
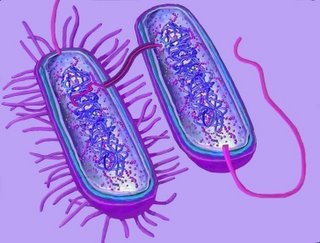
The middle diagram indicates passage of genetic material through a pilum from the F+ bacterium to a F- bacterium. This mechanism is under debate. (Compare with a 27,000 x magnification tem image tem conjugation E. coli)
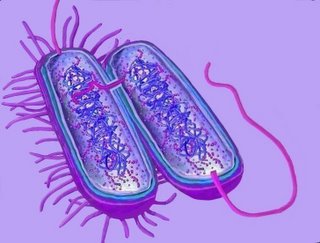
Transfer through the pili (above right) may not be a strictly accurate depiction of the actual transfer mechanism. The mechanism utilizes proteins coded by the tra or trb loci, and these may open a channel between the bacteria (bottom diagram). In this case, the pili are probably utilized to anchor and draw together the donor and recipient bacteria (top diagram). (Compare with conjugation in alga Conjugation of Spirogyra and a protist Conjugation of Paramecium.)
Once conjugation has been initiated by a mating signal, a complex of proteins called the relaxosome opens up one plasmid DNA strand at the origin of transfer, or oriT. The relaxosome system of the F-plasmid system comprises proteins TraI, TraY, TraM, and a protein that functions as the integrated host factor, IHF. The transferred strand – the T-strand – is unwound from the duplex before transfer into the recipient bacterium in a 5'-terminus to 3'-terminus direction. The remaining strand is replicated. Replication in concert with conjugation is termed conjugative replication, and is similar to the rolling circle replication of lambda phage. Replication may also occur independent of conjugative action – this is vegetative replication, beginning at the oriV.
Table Mechanisms of Biological Evolution : Gene Regulation in E.coli :









































Blue terms hyperlink to explanatory items. Linked items can also be found by way of the 'Links to this post' list at the base of some posts (once Blogger catches up!). Use the "back" function to return to the departure item.
Items occur within Sections. When visiting an item, the site title changes to purple – click on the title or “Home” to return to the main page. Topics are listed in the Site Map (click on arrow at top of sidebar). The site is searchable – once Blogger catches up – by way of the 'Search this blog' window at upper left.
When the number before the “Guide-Glossary” link (below each item) is greater than 0, the link provides a glossary of terms. Displayed as a pop-up when reading within a Section, or as sub-script when visiting an Item.
Allopatric speciation occurs when a geographical barrier sub-divides a parent species, resulting in geographic and reproductive isolation such that the descendent species can no longer interbreed upon removal of the barrier.
Anagenesis differs from cladogenesis in that one species progressively transforms into a replacement species when sufficient gene mutations fix in the descendant population. At this point, the ancestral species has become extinct. This mechanism is distinct from the increase in numbers of species generated by cladogenetic branching events.
Cladogenesis is the mechanism of speciation in which one or more lineages (clades) arise from an ancestral line. Such speciation events increase the variety of plants or animals through branching of the phylogenetic tree. Cladogenesis is differentiated from anagenesis, which is the in toto replacement of one species by an anatomically distinct species.
Monophyletic taxon or clade: an accurate grouping of only (opp. polyphyletic) and all (opp. paraphyletic) descendents of a shared common ancestor. A monopyletic group is genetically homogeneous and reflects evolutionary relationships.
Paraphyletic taxon or clade: a monophyletic group that excludes one or more discrete groups descended from the most recent common ancestral species of the entire group. Other descendent species of the most recent common ancestor have been excluded from the paraphyletic taxon, usually because of morphologic distinctiveness.
Phenetic system: groupings of organisms based on mutual similarity of phenotypic (physical and chemical) characteristics. Phenetic groupings may or may not correlate with evolutionary relationships.
Phylogenetic system: groups organisms based on shared evolutionary heritage. DNA and RNA sequencing techniques are considered to give the most meaningful phylogenies.
Phylogenetic separation into evolutionary relationships (clades), based on comparison of genomes is likely to supplant phenotypical (phenetic) taxonomies of the prokaryotes.
Peripatry (paripatry) is a subset of allopatry in which an isolated group has a smaller population than the parent group. Ernst Mayr introduced the term. Peripatric speciation occurs when the smaller sub-group of a species enters a novel niche within the range of the parent species, becoming geographically and reproductively isolated. Peripatric speciation (paripatric) is distinguished from allopatric speciation by the smaller size of the isolate group, and from sympatric speciation, which involves no barrier to breeding.
Polyphyletic taxon: opposite to monophyletic taxon: A polyphyletic group is mistakenly or improperly erected on the basis of homoplasy.—characteristics that have arisen despite not sharing a common ancestor. Homoplasy arises because of convergent evolution, parallelism, evolutionary reversals, horizontal gene transfer, or gene duplications. Polyphyletic taxa are genetically heterogeneous because members do not share a common ancestor.
Neontology is a branch of biology that emphasizes the study of modern biota (living or recent organisms) rather than fossilized organisms (paleontology).
Numerical Taxonomies are a common approach to phenetic taxonomy that employ a number of phenotypic characteristics to generate similarity coefficients that may be mapped in dendrograms. Groupings based on numerical taxonomy may or may not correlate with evolutionary relationships.
Taxonomies aim to group organisms according to shared characteristics against the background of biological diversity.
Sympatry involves no geographical separation of sub-populations of individuals. Sympatric speciation events occur most often in plants by the mechanism of polyploidy in which the number of chromosomes is doubled or tripled. John Maynard Smith proposed a model called disruptive speciation, in which homozygotes might have greater fitness than heterozygotes under some environmental conditions.